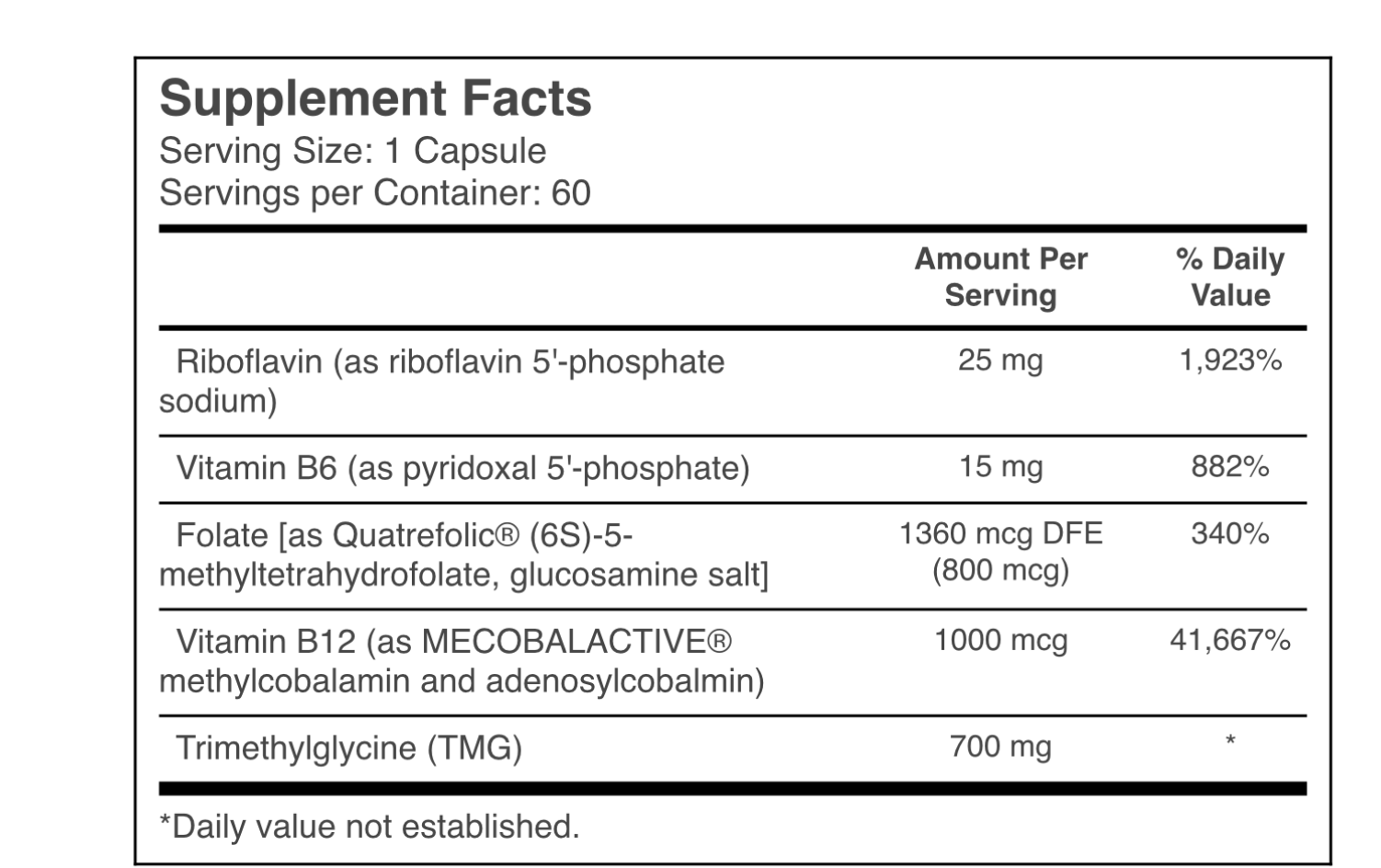
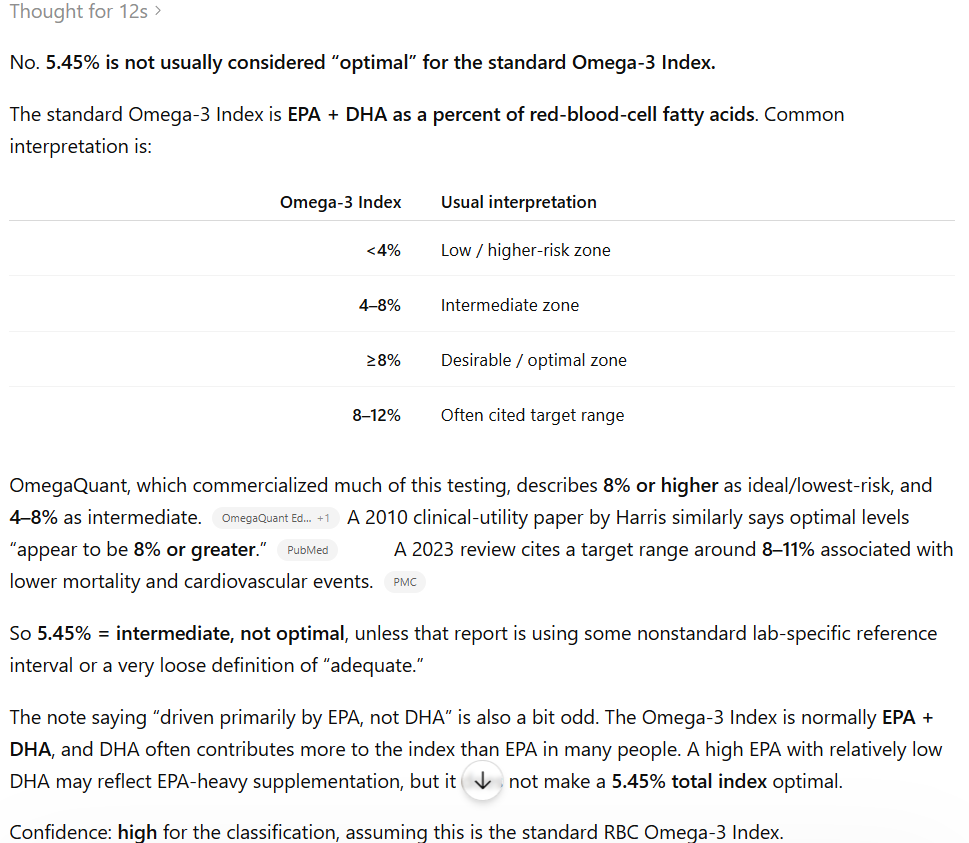
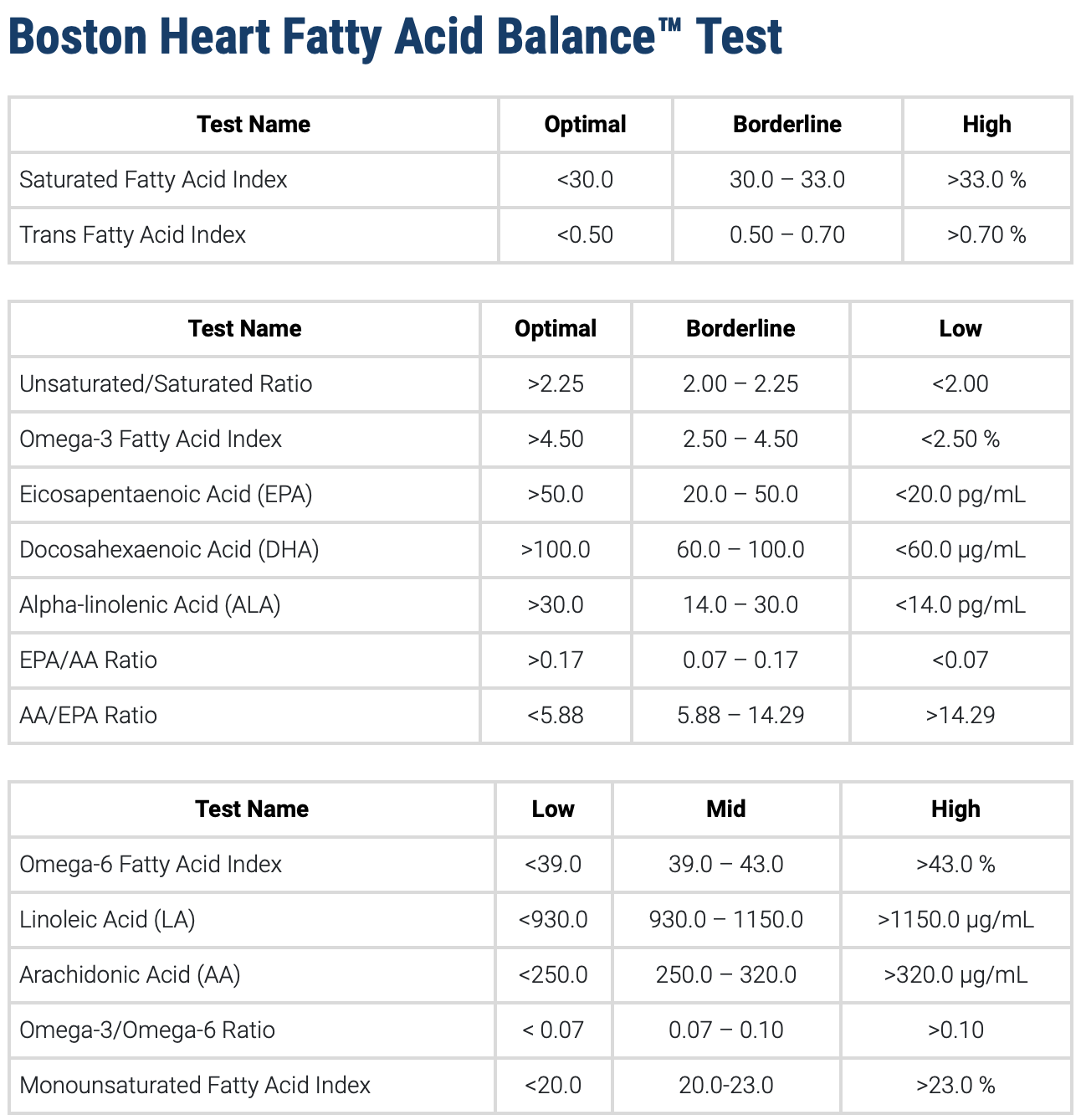
I’m taking the usual stuff for MTHFR C677T (heterozygous) without any success. I also tested for all the related B vitamins from B2 to B12 and they are all normal or even high.
Thanks to those pathway deep dives I now know that I have a severe NRF2 / glutathione bottleneck which by itself is enough to get high homocysteine.
I’ve started taking sulforaphane (Avmacol ES) for the NRF2 / glutathione bottleneck so we’ll see if that helps.
Here is Gemini 3 thinking explanation of that:
The short answer is yes. A bottleneck in the NRF2 pathway or a deficiency in glutathione (GSH) can lead to elevated homocysteine levels, even when your B-vitamin status and GGT (Gamma-Glutamyl Transferase) levels appear optimal.
This happens because the body prioritizes oxidative stress management over the methylation cycle. When glutathione is depleted or NRF2 is under-activated, the body diverts sulfur-containing precursors toward antioxidant production, creating a “metabolic pull” that can trap homocysteine.
1. The “Transsulfuration Pull” Mechanism
Homocysteine sits at a critical junction in your metabolism. It can either be recycled back into methionine (via B12 and Folate) or sent down the transsulfuration pathway to create cysteine, the rate-limiting precursor for glutathione.
-
The Bottleneck: If NRF2 is not sufficiently upregulating the enzymes required for glutathione synthesis, or if there is high oxidative stress, the body “calls” for more cysteine.
-
The Result: Homocysteine is diverted heavily into the transsulfuration pathway. If the downstream synthesis of glutathione is slow (the NRF2 bottleneck), homocysteine can “back up” because the cycle isn’t clearing it fast enough, or the body is intentionally keeping homocysteine production high to meet the perceived demand for antioxidants.
2. Why Low GGT and High B-Vitamins Don’t Rule This Out
It is common to assume that if B-vitamins are “high,” methylation should be perfect. However, in this specific context:
-
B-Vitamin Paradox: Having high serum levels of B12 or Folate does not mean they are being utilized efficiently at the cellular level. If the transsulfuration pathway is being heavily “pulled” due to glutathione demand, the methylation side (where those B-vitamins work) may become secondary.
-
The GGT Misconception: While high GGT is a marker of oxidative stress, very low GGT can actually be a sign of glutathione depletion. GGT is responsible for breaking down extracellular glutathione so the amino acids can be brought back into the cell. If there isn’t enough glutathione to “recycle,” GGT activity may drop significantly.
-
The NRF2 Factor: NRF2 controls the enzyme GCLC (Glutamate-Cysteine Ligase), which is the first step in making glutathione. If NRF2 is sluggish, you cannot convert cysteine into glutathione efficiently, leading to a build-up of the intermediates—including homocysteine.
3. The Role of Cystathionine Beta-Synthase (CBS)
In an NRF2/Glutathione bottleneck, the enzyme CBS is often up-regulated by oxidative stress. This enzyme pulls homocysteine out of the methylation cycle.
If your NRF2 response isn’t finishing the job (turning that cysteine into glutathione), you end up in a state of “high demand, low output.” The body continues to feed the transsulfuration pathway to fix the redox imbalance, keeping homocysteine elevated as it waits for the bottleneck to clear.
Summary Table: Why the Bottleneck Occurs
| Factor |
Status |
Impact on Homocysteine |
| NRF2 Activity |
Low/Sluggish |
Fails to trigger glutathione synthesis enzymes, causing metabolic backup. |
| GGT |
Low |
May indicate a lack of glutathione available for recycling/transport. |
| B-Vitamins |
High |
Shows the “machinery” is fueled, but the “priority” is shifted to antioxidants. |
| Oxidative Stress |
High |
Acts as a signal to keep diverting homocysteine away from methylation. |
To address this, the focus usually shifts from “more B-vitamins” to NRF2 activators (like sulforaphane) or direct glutathione support to signal the body that the oxidative crisis is over, allowing homocysteine to return to its normal recycling patterns.