A re-run of the previous prompt with a focus on 100X:
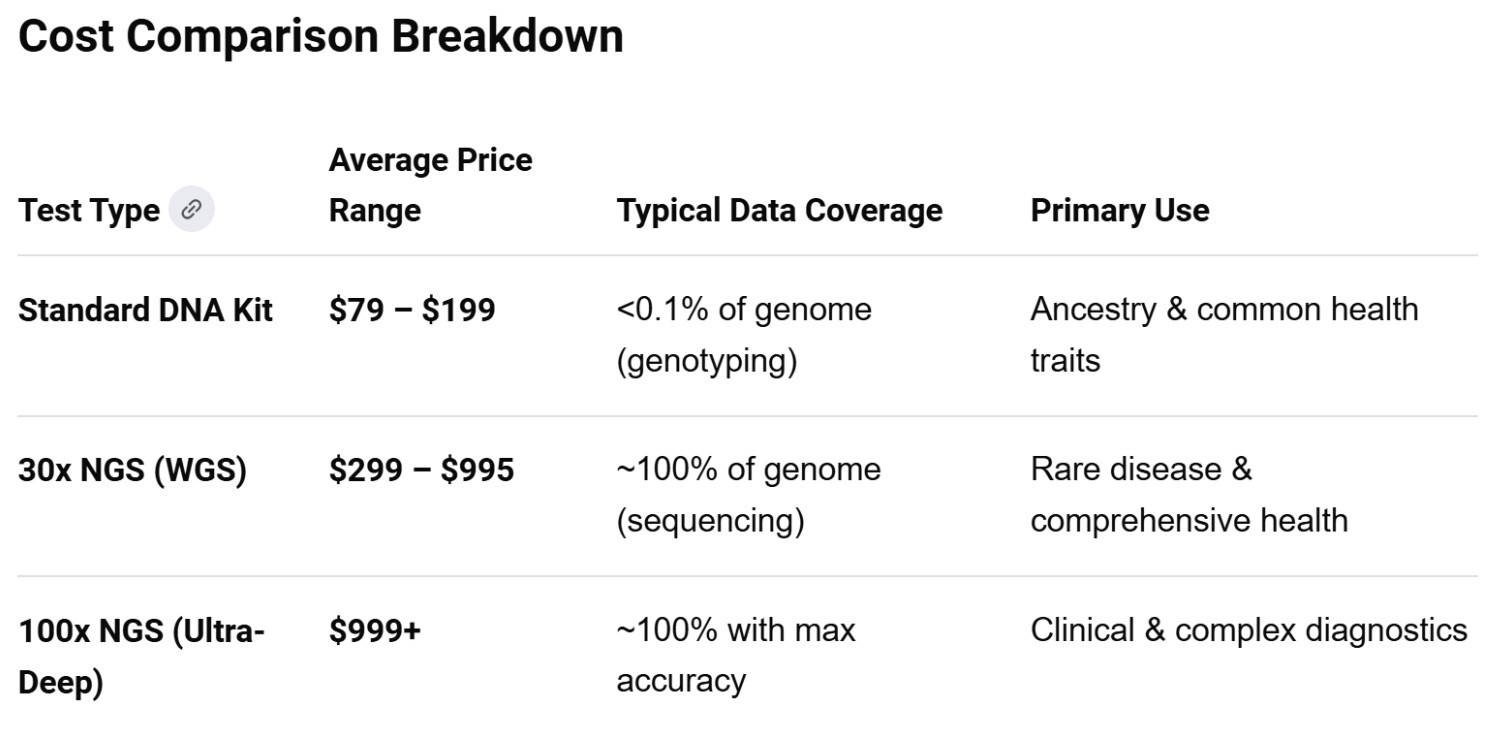
As of May 2026, the market for 100X Whole Genome Sequencing (WGS) has shifted significantly following the consolidation of several early pioneers (e.g., Nebula Genomics transitioning into DNA Complete). 100X sequencing, or “Ultra-Deep” sequencing, provides a higher signal-to-noise ratio than the 30X clinical standard, making it the preferred choice for detecting low-frequency somatic variants and mosaicism.
The following providers allow direct consumer purchase, provide raw VCF and CRAM files, and do not mandate a recurring subscription for data access.
Top 10 Lowest-Cost 100X Whole Genome Sequencing Sources (May 2026)
| Rank | Product/Brand Name | Vendor | Total Price (USD) |
|---|---|---|---|
| 1 | WGS Elite (100X WGS) | DNA Complete | $995 |
| 2 | WGS 100X Deep Sequencing Bundle | Sequencing.com | $1,099 |
| 3 | Whole GenomeZ (100X Coverage) | Dante Labs | $1,199 |
| 4 | WGS Custom (300 Gb Data Allocation) | YSEQ | $1,250 |
| 5 | Ultra-Deep 100X WGS | BioAro | $1,499 |
| 6 | Variant-Plus (100X Research WGS) | Macrogen | $1,650 |
| 7 | Personal Genome 100X Premium | Veritas Intercontinental | $1,850 |
| 8 | DeepSeq WGS Research Protocol | Psomagen | $1,900 |
| 9 | Clinical 100X Whole Genome | HiFiGenomics | $2,100 |
| 10 | Platinum 100X Research WGS | Illumina (Partner Network) | $2,500 |
Shipping and Logistics Summary
- DNA Complete: Currently offers a promotional free shipping rate within the Continental USA. International kits involve a flat $45 fee for delivery and return logistics.
- Sequencing.com: Domestic shipping is $19.99. For the 100X “Elite” bundle, return shipping is typically included via a prepaid label in the box.
- YSEQ: Shipping is calculated based on current rates from Germany. Expect roughly $25–$35 for the kit; the customer is responsible for return postage to their laboratory in Berlin.
- Dante Labs: Includes free global shipping, but users should note that turnaround times for 100X processing can exceed 12 weeks during high-volume periods.
- BioAro: Offers guaranteed turnaround times (under 4 weeks), but shipping costs vary by region, averaging $30 for North American customers.
Scholarly Analysis & Technical Considerations
The primary scholarly debate surrounding 100X WGS is whether the incremental benefit of increased depth justifies the $600–$800 premium over standard 30X sequencing.
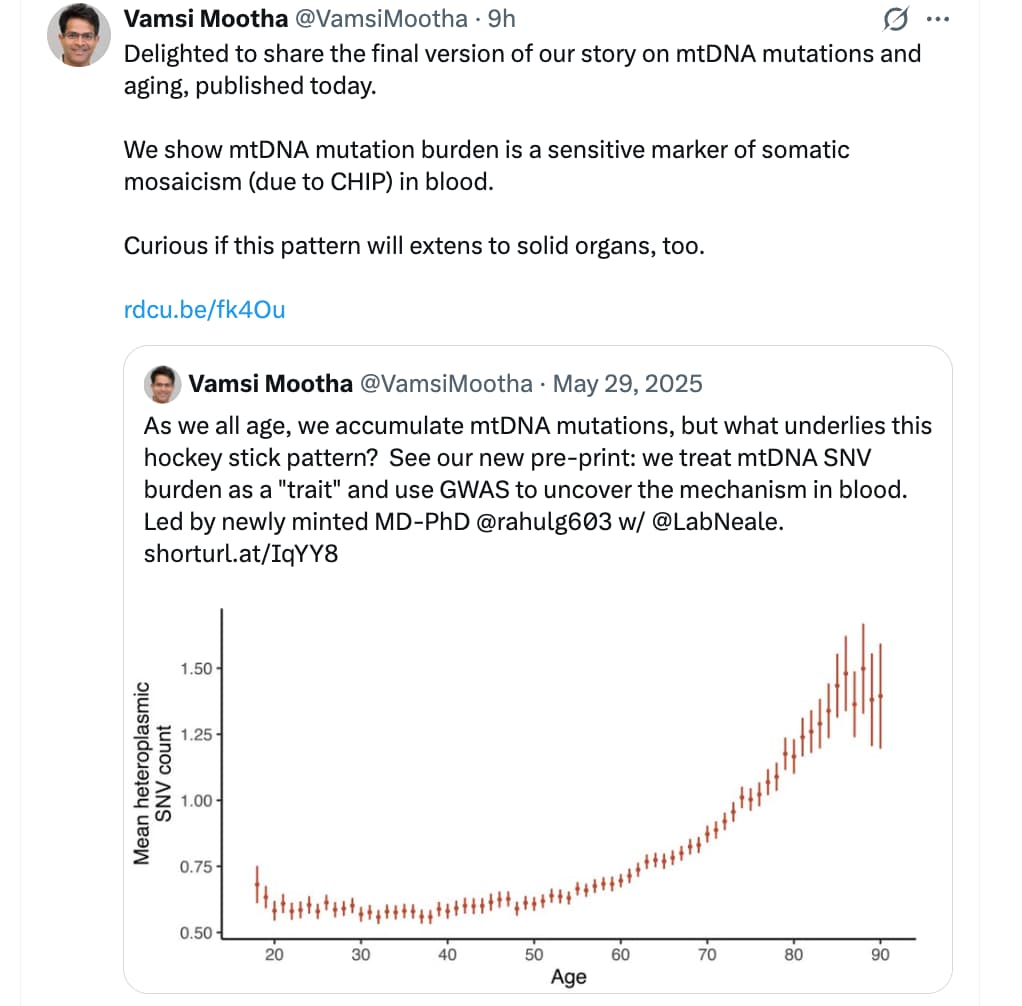
- Detection Thresholds: 100X depth is technically required for high-confidence identification of “mosaic” mutations (mutations occurring in only a fraction of cells), which are increasingly studied in the context of neurodegenerative diseases and longevity science.
- File Management: A 100X CRAM file typically exceeds 150GB. A knowledge gap exists in the consumer market regarding the necessary local hardware for processing such files; standard consumer-grade laptops often lack the RAM required for local alignment or variant browsing.
- No-Subscription Access: While the providers above allow one-time raw data downloads, several (DNA Complete, Sequencing.com) host “Report Marketplaces.” Users should distinguish between the sequencing data (which you own) and the interpretative reports (which may require separate fees).
Note on Reliability: Data for smaller providers (e.g., Full Genomes Corp) has been excluded due to inconsistent service availability and “Out of Business” flags in recent 2026 market audits. Always verify that the lab utilizes CAP/CLIA-certified pipelines if you intend to share the VCF with a medical professional for clinical intervention.