This one should be of interest for a lot of people as it shows who is at risk of of various side effects. For instance lipid and glucose dysregulation (like me for both! ![]() )
)
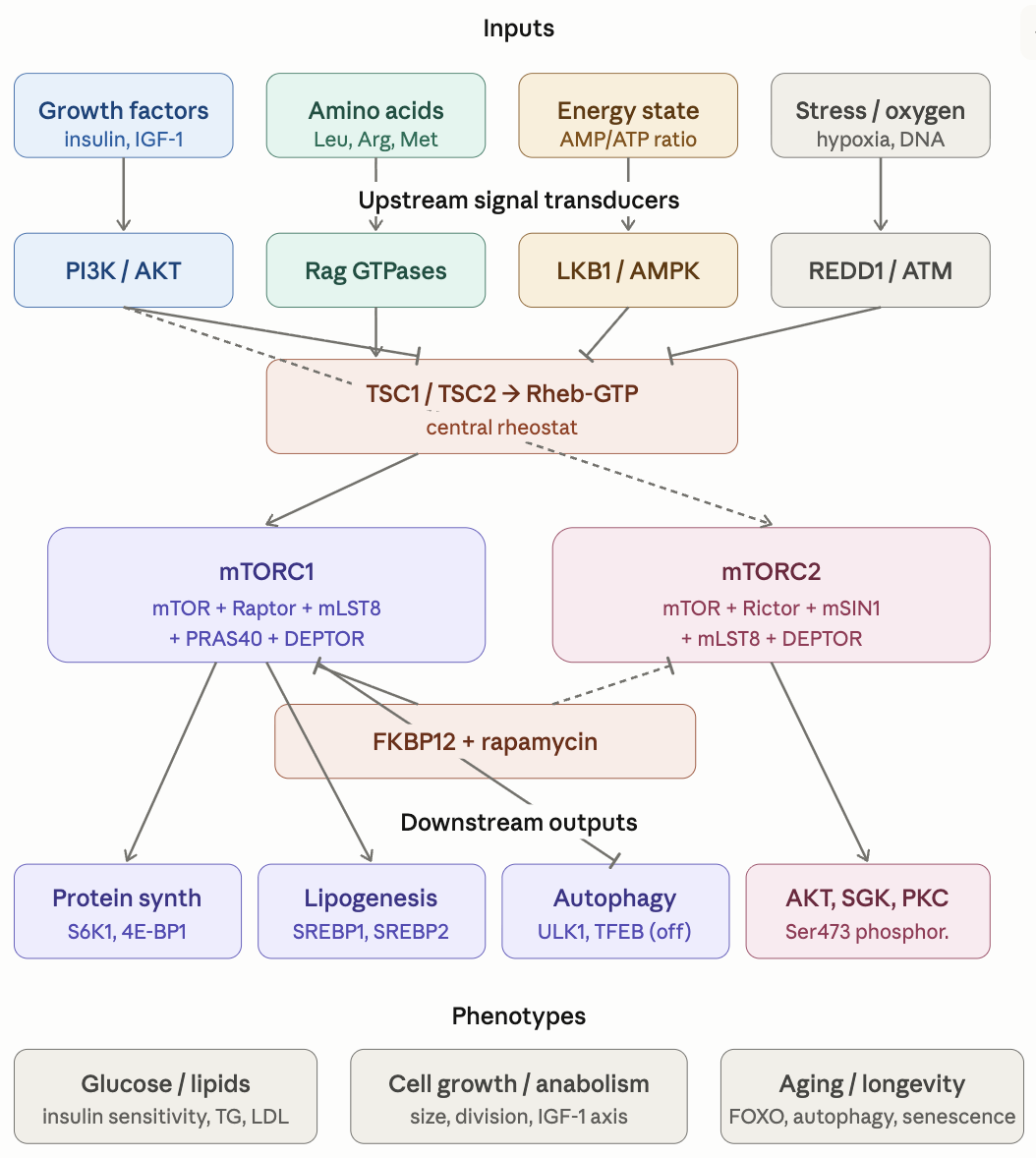
I’m continuing my deep dives into the genetic pathways to get actionable insights as the previous ones have been incredible precise and useful. This time I’m looking at rapamycin genetic pathways.
Here is the general description of the pathways and their variants. I will put the finding about my own genome below it as an example of what useful and actionable insights you can get.
Rapamycin_mTOR_Genetic_Pathway_Reference.pdf (507.8 KB)
The pdf report above is valid for everybody but here is the summary of the findings I get when I apply it to my own genome. (The full detailed report is 21 pages long.)
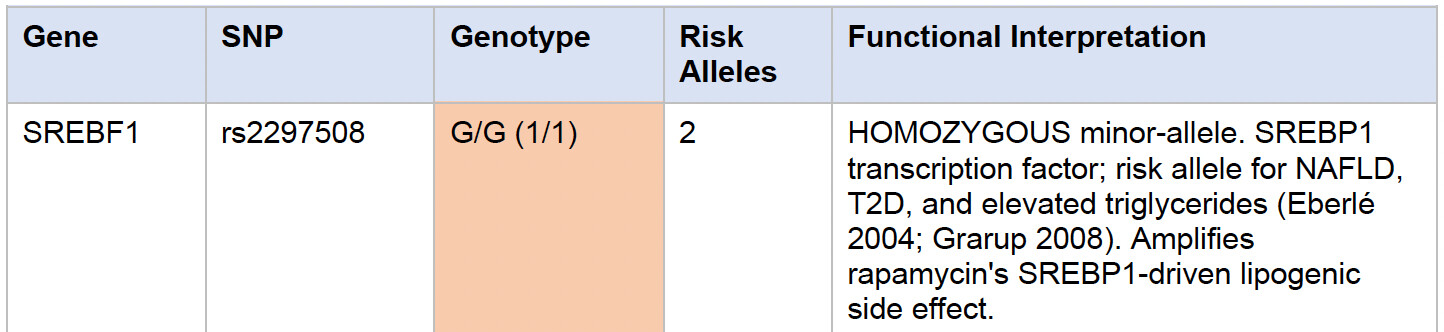
Lipids
The homozygous SREBF1 rs2297508 G/G is a moderately well-replicated NAFLD/T2D/triglyceride risk variant; on rapamycin (which raises TG and LDL via SREBP1/2 activation), this genotype amplifies the lipogenic response.
Glucose
The TBC1D4 homozygous status modulates the AKT → TBC1D4 → GLUT4 translocation axis — exactly the chain that mTORC2 inhibition by chronic rapamycin disrupts (Lamming 2012). The PCK1 het pushes additionally on the gluconeogenic output side. The IRS1 protective C/C is the partial offset. Net interpretation: the patient is genetically positioned in the upper half of the population distribution for rapamycin-induced fasting hyperglycemia. The observed clinical phenotype is mechanistically expected. This does not contraindicate continued rapamycin use; it argues for (a) trough-glucose monitoring around dosing days, (b) confidence in the AMPK-active co-therapy (imeglimin) and GLP-1 RA (tirzepatide)