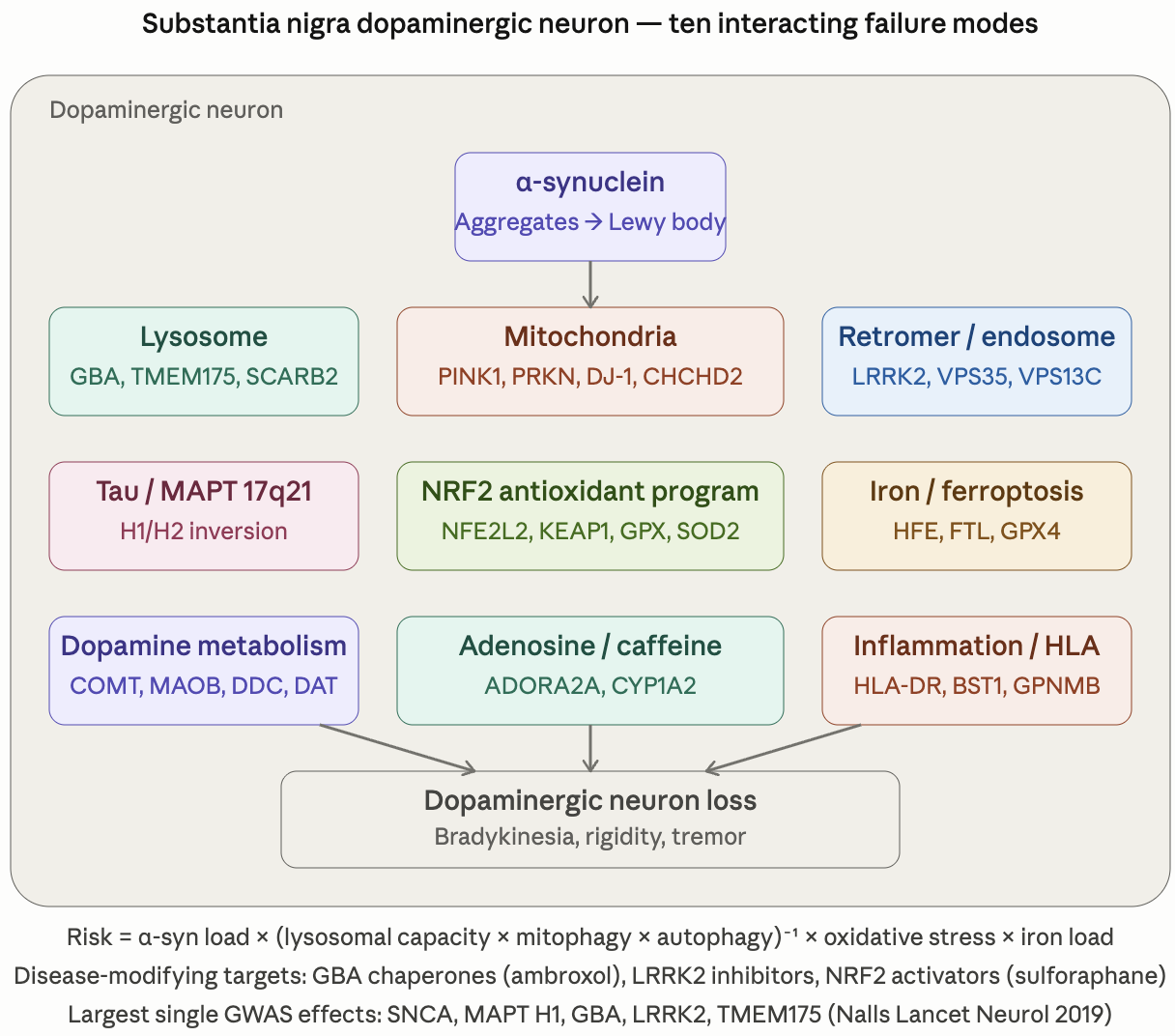
I’m continuing my deep dives into the genetic pathways to get actionable insights as the previous ones have been incredible precise and useful. This time I’m looking at the Parkinson’s related genetic pathways.
Here is the general description of the pathways and their variants. I will put the finding about my own genome below it as an example of what useful and actionable insights you can get.
Parkinson_Disease_Genetic_Pathway_Reference.pdf (681.0 KB)
The pdf report above is valid for everybody but here is the summary of the findings I get when I apply it to my own genome.
Parkinson’s Disease Genetic Report — Top actionable findings
Cumulative integrated PD risk approximately at population baseline to mildly elevated (estimated common-variant PRS ~50th–65th percentile, ~1.0–1.2× population baseline lifetime risk). No high-impact rare/Mendelian PD variants detected. The dominant risk axis is the recurring NRF2 / glutathione keystone (already actively addressed). The dominant favorable feature is the apparent MAPT H2/H2 protective haplotype (PSP/CBD risk substantially below population baseline). Existing supplement and medication regimen is well-aligned with the genetic profile.
1. NRF2 keystone confirmed across THREE reports — sulforaphane stack is mechanistically optimal (highest priority)
NFE2L2 rs6721961 G/G (homozygous risk) + rs2706110 C/C (homozygous risk) + rs35652124 C/T (het). Same rs6721961 G/G called in Inflammation/Immune and Glycation reports. Plus GSTP1 105V/V + 114V/V double homozygous (glutathione conjugation impairment) and NQO1 *2 heterozygous. Action: continue Avmacol Extra Strength BID + NACET 100 mg AM / 200 mg PM + glycine 6 g. Form and dose are correct — no change. Independently supported by the PD literature now (Bento-Pereira Med Res Rev 2021; Jazwa Antioxid Redox Signal 2011).
2. MAPT H2/H2 protective — largest favorable finding
All six independent H1-tagging MAPT SNPs (rs1052553, rs17649553, rs8070723, rs393152, rs242557, rs2435207) PLUS both flanking 17q21 inversion tags (KANSL1 rs17563986, CRHR1 rs17689882) are homozygous reference — most parsimoniously H2/H2 (population frequency ~4% in Europeans). Per-allele PSP OR ~0.18, PD OR ~0.7. Action: informational only; no intervention. Optional: confirm with phased haplotype analysis (shapeit) if preferred — the chance of all six H1 tags being hom-ref by chance in an H1-carrier is vanishingly small, but formal phasing is reassuring.
3. All Mendelian / high-impact PD variants ABSENT — major favorable
Hom-ref for: LRRK2 G2019S (rs34637584), LRRK2 R1441C (rs35870237), LRRK2 non-coding rs76904798, VPS35 D620N (rs188286943), GBA1 N370S/L444P/E326K/T369M, SNCA A53T/A30P/E46K/H50Q/G51D, PINK1 Q456X/G411S, PRKN R275W/D394N coding, CHCHD2 T61I, SMPD1 L302P. Action: none. Caveat: SNCA duplication/triplication and PRKN exonic CNVs are not detectable from short-read SNP VCF; not clinically indicated to pursue absent family history of early-onset PD.
4. SNCA risk haplotype heterozygous (not homozygous) — modest
SNCA rs356182 HET + rs356219 HET + rs2737029 HET (one risk allele each at three tag SNPs in linkage disequilibrium). Top common-variant PD GWAS signal but at heterozygous level (per-allele OR ~1.32). Action: none specific. Rapamycin (autophagy/α-syn clearance) and doxycycline (α-syn aggregation modulation) in current regimen are mechanistically aligned. Note: the heterozygous rs356182 G/A genotype does not predict tremor-predominant phenotype (that finding was specific to G/G homozygotes; Cooper Mov Disord 2017).
5. Pesticide G×E susceptibility profile — minimize modifiable exposure
GSTP1 105V/V (rs1695) + 114V/V (rs1138272) double homozygous + NAT2 *5 slow-acetylator homozygous (rs1801280 C/C) + SLC11A2 (DMT1 iron/Mn) homozygous (rs422982) = textbook Menegon-Lancet-1998 pesticide-PD G×E susceptibility configuration. Action: test private well water if applicable; rinse non-organic produce; avoid organophosphate lawn treatments and welding/manganese fume exposure. No supplement change required (selenium 100 µg/d in Momentous Multi already supports GPX4).
6. Coffee/caffeine — continue moderate intake (1–4 cups/d)
ADORA2A partial-protective haplotype: rs5751876 HET + rs5760423 HET + rs2298383 HET (the strong rs3032740 functional variant is hom-ref, so A2A receptor density is at baseline). CYP1A2 *1A/*1F (rs762551) C/A heterozygous = intermediate metabolizer. Coffee–PD inverse association is the most replicated environmental neuroprotection signal (Hernán Ann Neurol 2002; Ascherio Lancet Neurol 2016; Grade A). Action: continue current intake.