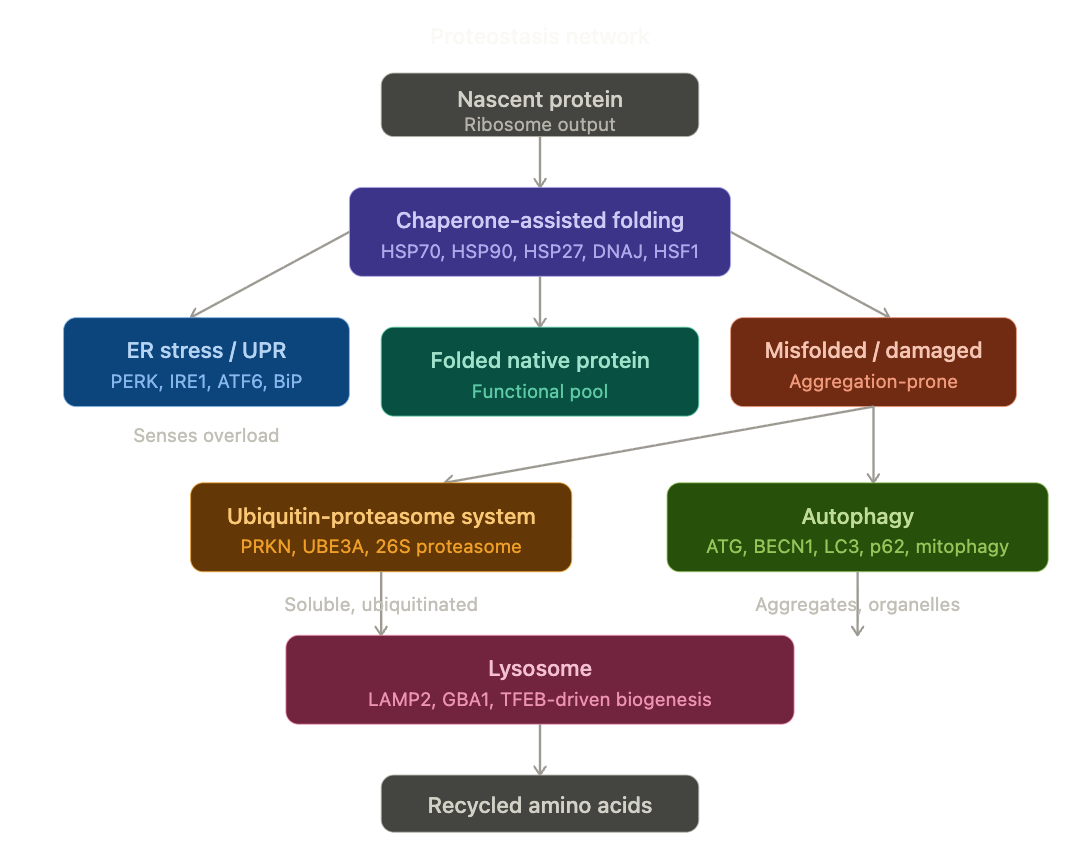
I’m continuing my deep dives into the genetic pathways to get actionable insights as the previous ones have been incredible precise and useful. This time I’m looking at the Autophagy / Proteostasis / Unfolded Protein Response related genetic pathways.
Here is the general description of the pathways and their variants. I will put the finding about my own genome below it as an example of what useful and actionable insights you can get.
Proteostasis_Genetic_Pathway_Reference.pdf (485.7 KB)
The pdf report above is valid for everybody but here is the summary of the findings I get when I apply it to my own genome.
Top findings (highest evidence + highest expected impact)
1. Homozygous EIF2AK3 PERK haplotype B — rs7571971 T/C 1/1, rs13045 T/C 1/1, rs1805165 C/A 1/1 (DP 72-97, GQ=99 throughout). All three coding nonsynonymous SNPs PLUS the GWAS-significant intronic tag are homozygous on the same haplotype block. Maximum-load PERK-B configuration (~8% population frequency homozygous). Reduced stress-induced eIF2α phosphorylation; increased ER-stress sensitivity in tissues with high secretory load (β-cells, hepatocytes, osteoblasts). Validated associations: PSP risk OR 1.4-1.6 per allele (Höglinger 2011 Nat Genet, p=3.2×10⁻¹³); reduced BMD (Liu 2012 JBMR); growth hormone deficiency association (Yuan 2018 HMG); HIV-NCI risk (CHARTER 2022). Does NOT cause Wolcott-Rallison syndrome (which requires biallelic LoF, not the missense haplotype). The single most consequential new finding of this report.
2. Homozygous ATG16L1 T300A — rs2241880 A/G 1/1 (DP=95, GQ=99). Threonine→alanine substitution at residue 300 creates a caspase-3 cleavage site. Under any caspase-3-activating insult (oxidative stress, inflammatory signaling, infection, ER stress) ATG16L1 protein is preferentially degraded. The defect is selective: bulk macroautophagy (mTOR-regulated, rapamycin-responsive) is preserved; the impaired arms are xenophagy, ER-phagy, and unconventional autophagy. Crohn’s risk OR 1.23 in Caucasians (Hampe 2007 Nat Genet, replicated 100+ studies). Among the most replicated common autophagy variants in human genetics.
3. Homozygous NFE2L2 −617 risk genotype — rs6721961 T/G 1/1 on forward strand = A/A on coding strand (DP=76, GQ=99). Third independent confirmation across reports (PD, Glycation, Endurance). Reduces NRF2 promoter activity by approximately 30-60%. Proteostasis-specific consequence: blunts inducible expression of p62/SQSTM1 (the major aggrephagy receptor), GCLC + GCLM (glutathione synthesis subunits), HMOX1, and NQO1. The dominant cross-report keystone finding.
4. Homozygous mTOR triplet — rs2295080 G/T 1/1, rs1057079 C/T 1/1, rs11121704 C/T 1/1 (DP 50-103, GQ=99 throughout). Promoter and 3’UTR variants associated with elevated mTOR expression and wet AMD risk (Wei 2020). Combined with AKT1 rs1130214 1/1 (cross-ref IGF1 report — strong AKT phosphorylation suppresses FOXO3) plus homozygous IGF1 alleles (cross-ref IGF1) defines a high-mTOR / AKT-strong signaling configuration.
5. FOXO3 longevity haplotype heterozygous — rs2802292 G/T 0/1, rs2802288 A/G 0/1 (DP 73-87, GQ=99). Cross-confirms the IGF1 report’s six-SNP heterozygous longevity haplotype including both fine-mapped causal variants (rs12206094 + rs4946935). The G allele creates an HSF1-binding enhancer in intron 2 that drives stress-induced FOXO3 transcription independently of AKT phosphorylation (Donlon NAR 2018). Per-allele longevity effect ~1.4× probability of reaching age 95 (Willcox PNAS 2008; Bao npj Aging 2024). The single most important favorable genetic finding in this proteostasis profile.
6. HSP70 cluster all homozygous reference — six common functional SNPs in HSPA1A/HSPA1B/HSPA1L (rs1043618, rs1008438, rs2227956, rs2075800, rs1061581, rs2763979) returned no variant calls at 60× coverage, consistent with full hom-ref across the entire chr6p21.3 cluster. Full inducible cytosolic chaperone capacity preserved — a critical compensatory backstop for the homozygous PERK-B finding above.
7. All Mendelian / high-penetrance proteostasis variants ABSENT — GBA1 N370S/L444P/E326K/T369M, VPS35 D620N, LRRK2 G2019S/rs76904798, PINK1, PRKN, PARK7, DNAJC13 N855S, HSPB1 CMT2F missense, OPTN ALS-FTD missense, SQSTM1 PDB/ALS missense, LAMP2 Danon-disease LoF — all clean. Cross-confirms PD and Dementia reports.
8. ATG7 V471A and P426L absent — rs36117895 and rs143545741 both not in VCF. The NAFLD-progression alleles (Iorio Hepatology 2024) are absent; ATG7 E1-like activity preserved at the protein level. Two intronic ATG7 heterozygotes (rs7635838 HDL-tag, rs2447607 BP-tag) are functionally minor. The autophagy elongation arm has the ATG16L1 defect but NOT the ATG7-side defect — partial rather than compounding lesion.
Top priority interventions
1. CONTINUE rapamycin 12 mg every 3 weeks. The single most genotype-aligned medication in the regimen. Directly counters the homozygous mTOR triplet AND the homozygous AKT1 (cross-ref IGF1), removes the AKT-driven FOXO3 suppression that prevents the heterozygous longevity allele from being fully expressed, AND pushes flux through the canonical autophagy pathway that is preserved (compensating for ATG16L1 T300A). Also dephosphorylates TFEB → lysosomal-biogenesis induction, augmenting the user’s intact lysosomal compartment. Rapamycin has four distinct mechanistic alignments with this genotype. No dose change recommended.
2. CONTINUE Avmacol Extra Strength 4 caps/day total. Already optimized to ~40-60 mg sulforaphane equivalents per the Endurance report. Sulforaphane bypasses the rs6721961 promoter deficit at the KEAP1 cysteine-modification step, sustaining longer NRF2 protein half-life once translated. Proteostasis-specific bonus: augments the inducible p62 pool, partially compensating for ATG16L1 T300A.
3. CONSIDER ADDING heat-stress exposure (sauna 20-30 min at 80-90°C, 3-5×/week if accessible). The FOXO3 longevity haplotype’s HSF1-binding enhancer is specifically engaged by heat-shock — this is the single most direct way to activate the heterozygous longevity allele and partially override AKT-driven FOXO3 suppression. Mechanistic backing: Donlon NAR 2018 demonstrated HSF1 occupancy at the rs2802292 G-allele enhancer is induced by heat-shock; observational human data: Laukkanen JAMA Intern Med 2015 (Finnish sauna study, dose-response cardiovascular and all-cause mortality reduction); Brunt J Physiol 2016/2018 (passive heat therapy improves endothelial function and arterial stiffness).
4. CONSIDER ADDING spermidine (1-3 mg/day; wheat germ extract or supplement). Direct autophagy inducer through inhibition of EP300 acetyltransferase (Pietrocola Cell Death Differ 2015), mTOR-independent. Provides a parallel autophagy push compatible with rapamycin and aligned with ATG16L1 T300A. Population-level observational data: Eisenberg Nat Med 2016 (dietary spermidine intake associated with reduced all-cause mortality); not yet RCT-validated.
5. MAINTAIN sustained metabolic-stress reduction via tirzepatide + empagliflozin + imeglimin. Chronic metabolic stress (glucose excursions, hepatic ER stress, mitochondrial ROS) is the major upstream caspase-3 trigger that makes ATG16L1 T300A clinically impactful. The user’s GLP-1/SGLT2/imeglimin regimen is genetically aligned for this. Should not be deprioritized; the current configuration is essentially optimal for this genotype.
Top contraindications and cautions
1. AVOID hydroxychloroquine and chloroquine (lysosomal alkalinizers) where alternatives exist. These suppress autophagy by raising lysosomal pH, more impactful given homozygous ATG16L1 T300A. Not currently indicated; flag if proposed for autoimmune indication.
2. AVOID chronic high-dose NSAIDs (sustained ibuprofen, naproxen at full doses) where alternatives exist. Chronic NSAID activation of caspase-3 in gut epithelium amplifies the ATG16L1 T300A defect. Aspirin 81 mg has minimal caspase-3 effect; continue.
3. AVOID sustained high-fructose intake and prolonged saturated-fat excess. Both directly induce hepatic ER stress, more impactful given homozygous PERK-B haplotype.
4. AMBROXOL (GBA1 chaperone) is NOT genetically indicated. All four GBA1 missense variants are absent (cross-ref PD report). Although ambroxol is sometimes promoted as a longevity/PD-prevention supplement, this user’s GBA1 is genetically clean and there is no rationale to add it.
5. CAUTION with bortezomib or other proteasome inhibitors if ever clinically proposed. Homozygous ATG16L1 T300A means less autophagy reserve to compensate for proteasome inhibition. Not currently indicated; flag for future clinical scenarios.