I’m continuing my deep dives into the genetic pathways to get actionable insights as the previous ones have been incredible precise and useful. This time I’m looking at the ADHD/ADD related genetic pathways.
Here is the general description of the pathways and their variants. I will put the finding about my own genome below it as an example of what useful and actionable insights you can get.
ADHD_Genetic_Pathway_Reference.pdf (566.6 KB)
The pdf report above is valid for everybody but here is the summary of the findings I get when I apply it to my own genome.
Executive Summary
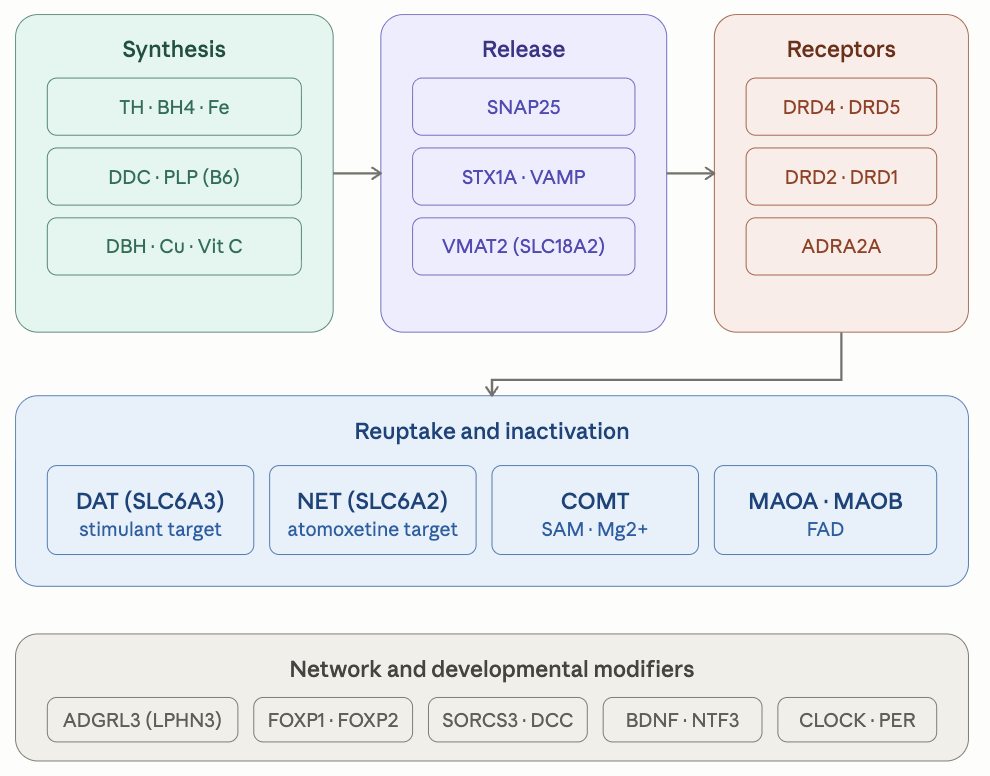
Analysis of 79 SNPs across ten functional categories of the ADHD genetic pathway identified 41 variants in the patient’s VCF. The genetic profile shows a striking convergence pattern: a heavily loaded adrenergic axis with multiple homozygous risk variants in ADRA2A and SLC6A2 (the noradrenergic transporter), a homozygous childhood-ADHD risk haplotype in ADGRL3 (latrophilin-3), a homozygous SNAP25 risk haplotype affecting vesicular release, and a uniformly fast catecholamine catabolism profile (COMT Val/Val from prior reports + MAOA hemizygous high-activity + MAOB hemizygous high-activity). Several common GWAS effector loci are homozygous (PCDH7, FOXP1, FOXP2). The dopamine receptor layer is largely favorable (DRD2 likely Glu/Glu from Strength report, DRD1 het, DRD3 het). The CYP2D6 pharmacogenomic locus, cross-referenced from the prior Pharmacogenomics report, supports likely normal-metabolizer status for atomoxetine.
This profile is internally coherent. The combined fast-clearance catabolism axis (COMT Val/Val + MAOA high + MAOB high + DBH heterozygous) predicts a modest tendency toward low-end-of-optimum prefrontal catecholamines under demanding cognitive load — the biological substrate that responds to NET-blocking pharmacology (atomoxetine, methylphenidate, viloxazine) and α2A agonists (guanfacine, clonidine). The ADRA2A risk-allele homozygous status independently predicts strong methylphenidate response (Liu et al. meta-analysis OR 3.08). The ADGRL3 haplotype is the largest single common-variant burden in this profile and is associated with childhood-onset, persistent-into-adulthood ADHD.
Primary Genetic Findings (homozygous risk burden)
- ADRA2A risk haplotype HOMOZYGOUS — rs1800544 (−1291 promoter, decoded), rs521674 (intronic tag), and rs553668 (3′UTR) all homozygous variant. ADHD inattention symptom association (Roman et al. 2003, 2006); favorable methylphenidate response prediction (Liu et al. 2021, OR 3.08).
- ADGRL3 / LPHN3 risk haplotype HOMOZYGOUS — rs6551665, rs1947274, rs1947275, and rs6858066 all homozygous variant. Latrophilin-3 four-SNP haplotype is the most replicated common-variant signal for childhood ADHD across Latin American, European, and meta-analytic samples (Bruxel et al., Mol Psychiatry 2021, PMID 32051549).
- SNAP25 risk haplotype HOMOZYGOUS — rs3746544 T/T plus rs1051312 C/C, the canonical 3′UTR T1065G/T1069C MnlI/DdeI haplotype (Liu et al. 2017, PMID 26941099). Predicts reduced presynaptic SNAP25 expression in the prefrontal cortex.
- SLC6A2 (NET) homozygous variant cluster — rs28386840 promoter region plus rs2242446 intronic tag, both homozygous. The norepinephrine transporter is the direct target of atomoxetine and a partial target of methylphenidate.
- MAOA hemizygous high-activity (rs6323 T) plus MAOB hemizygous high-activity (rs1799836 C) — male X-linked. Combined with COMT Val/Val (from prior reports), defines a fast catecholamine-catabolism profile.
- PCDH7, FOXP1, FOXP2 GWAS effector loci homozygous — rs1556812, rs7593947, rs12533005 — all hits from Demontis 2019/2023 GWAS at small per-allele effect sizes; cumulative polygenic burden meaningful.
- SPR (sepiapterin reductase) homozygous variant — rs1876487. SPR is a BH4-recycling enzyme; BH4 is the obligate cofactor for tyrosine hydroxylase (the rate-limiting catecholamine-synthesis enzyme) and TPH2.